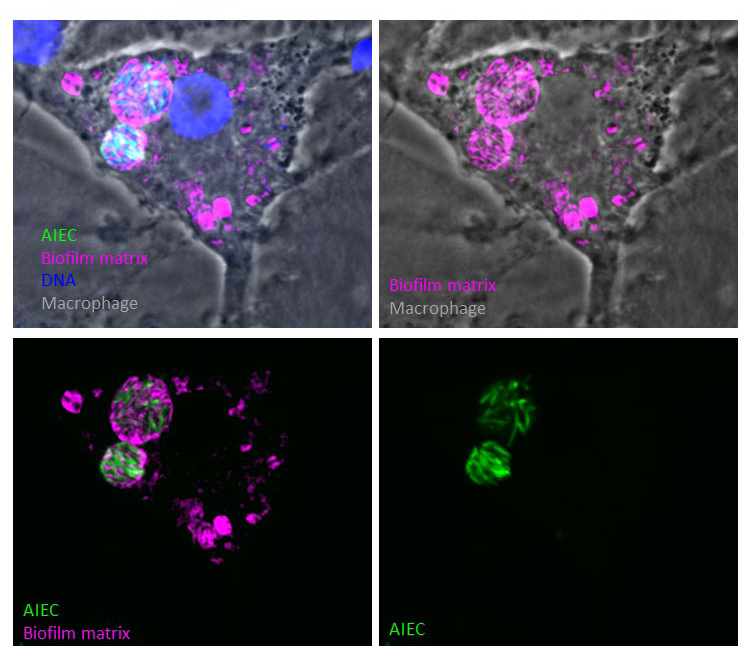
Our group studies the adaptation of bacteria to changes in their environments. Although bacteria are among the simplest living organisms, formed of a single cell, they display a huge adaptability to the presence of nutrients or toxic compounds in their environment. This adaptability allows them to survive periods of nutrient depletion, treatment with antibiotics or survival within their host in the context of an infection.
The first line of adaptation goes through the modulation of gene expression. This characteristic is known for decades. Since the pioneer work of Francois Jacob, Jacques Monod and their colleagues on the lactose operon in 1961, this field has been revolutionized by the development of genomic methods. In the laboratory, we use cell biology and genomic assays to analyze the expression of genes in response to various stimuli, the binding of proteins to multiple sites on the bacterial chromosome and even the 3D folding of the chromosome in various growing conditions.
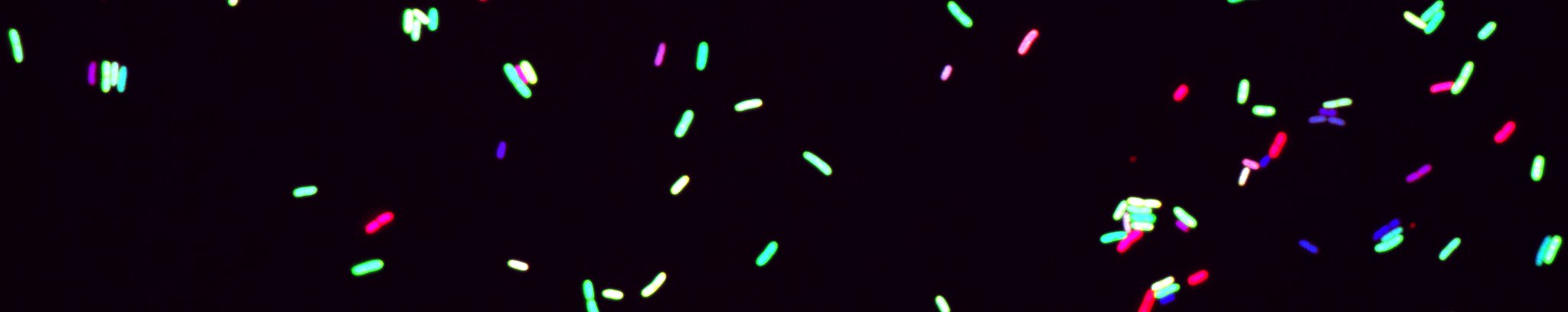
We observed that chromosome folding and genome expression are tightly connected (Lioy et al 2018, Planchenault et al 2020).We study various aspects of the interplay between chromosome folding and gene expression; we analyze of the role of the physics of the DNA molecule in transcription initiation and the role of transcription on long range DNA folding.
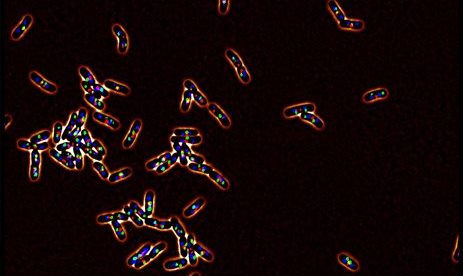
A second aspect of bacterial adaptation relies in modification of the growing regime and life style of the bacteria. In the presence of a stress bacteria can halt their cell cycle to limit the extent of the damages; in the harshest conditions, some bacteria may enter in a dormancy state, where metabolic activities are limited. This state confer them a high tolerance to antibiotic for example. In other conditions, bacteria cooperate to form communities, also called biofilm, that favor their adherence and tolerance to stresses.
In the recent years we have observed, with commensal Escherichia coli, that chemotherapeutic drugs such as Mitomycine C, Bleomycin or AZT severely change the cell cycle. This is linked to the sudden expression of dozen of genes from the SOS regulon. One protein from this regulon caught our attention, RecN. This protein belongs to the SMC family that also contains eukaryotic cohesins and condensins . When a particular type of DNA Damage repair is engaged, RecN is able to block chromosome segregation and to promote the regression of already segregated sister chromatids. RecN acts as a DNA damage specific cohesin, this is the first bacterial cohesin described (Vickridge et al 2017). In E. coli, sister chromatid cohesion is a transient step usually mediated by topological links between sister chromatids. Our group contributed to the discovery of this topological cohesion process (Lesterlin et al 2012) and characterized some aspect of its regulation (El Sayyed et al 2016). We studied the impact of the sister chromatid cohesion step on global chromosome folding in normal or pathological cell cycles (Conin et al 2022).